
Applied Conservation, Community Ecology, Foraging Interactions, Stable Isotope Biogeochemistry
University of California, San Diego, Division of Biological Sciences,
Ecology, Behavior, and Evolution Section


Among other things, I have studied the ecological factors driving trophic cascades in Sierra Nevada streams, demonstrated how invasive rats structure rocky intertidal ecosystems on the Aleutian Islands, Alaska via a cross community trophic cascade, tested for the presence/absence of invasive mammals in the San Juan Islands, and used stable isotope analysis to track vertebrate foraging ecology and movement patterns in Alaskan waters.




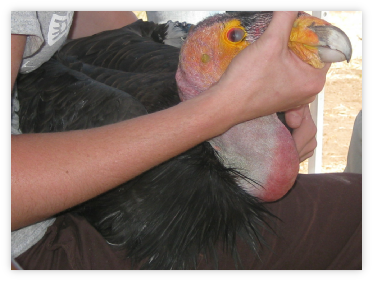
Marine derived contamination in California condor diets. In this collaborative research, we are using stable isotopes of nitrogen and carbon to determine the foraging ecology of California condors in order to assess the risks of ingesting marine derived pollutants present within marine mammal carcasses stranded on the central California coast. Photo by C. Kurle.
Estimating foraging patterns for northern fur seals in Alaska and California. We are using stable isotope analysis of whiskers from northern fur seal pups, juveniles, and adults to assess their foraging ecology and movement patterns during their non-breeding seasons. These projects are collaborations with scientists at NOAA’s NMML . Photo by A. Orr (NOAA/NMML).


Stable isotope discrimination factors for vertebrates. Stable isotope mixing models work best when the isotope discrimination factors (the isotopic difference between predator and prey tissues) are known. We are working on a variety of captive animals to determine these values. Photo by Ariana Ananda (UCSD undergraduate).
Sea turtle foraging ecology and movement patterns. In collaboration with scientists at NOAA’s SWFSC and Grupo Tortuguero de las Californias, we are extracting sea turtle growth lines from dead stranded turtles and analyzing their stable isotope values to determine ontogenetic shifts in sea turtle habitat use. Also in collaboration with NOAA, we are using compound specific stable isotope analysis to elucidate mechanisms driving a trophic dichotomy between leatherback turtles from different regions of the world. Photo by C. Turner Tomaszewicz (UCSD graduate student, Kurle lab)

Niche partitioning and impacts of foraging differences in rock iguana species on Caribbean islands. In collaboration with the San Diego Zoo Institute for Conservation Research and Earlham University, we are studying aspects of rock iguana foraging ecology to determine how diet drives size differences, population declines, and other aspects of iguana natural history. Photo by collaborator J. Iverson (Earlham University).

Maternal foraging strategies in Antarctic fur seals utilizing stable isotope analysis of tissues from their pups. In collaboration with NOAA’s Southwest Fisheries Science Center, we are analyzing the stable isotope values of fur seal pup vibrassae and fur to determine if they are appropriate proxies to estimate maternal diet and movement of Antarctic fur seals at Cape Shirreff, Livingston Island, Antarctica. Photo by collaborator M. Goebel (NOAA).

Quantifying niche partitioning in animals using stable isotope analysis and Bayesian
mixing models. We are using multiple mixing models to best determine the intraspecific
foraging niche differences in various animal species including American black bears
and invasive Norway rats to better inform conservation and management of species.
This project is in collaboration with Dr. Jack Hopkins, a post doc in the Kurle lab.
Rat ph![]() oto by S. Reisewitz, bear photo by J. Hopkins.
oto by S. Reisewitz, bear photo by J. Hopkins.
Utilizing long-term tissue archives to reconstruct oceanographic patterns in relation to potential environmental variability. In collaboration with CalCOFI and NOAA’s NMML and SWFSC, we are utilizing compound specific stable isotope analysis of amino acids from multiple tissue archives from northern fur seals, sea turtles, and plankton, to reconstruct changes in primary production, nitrogen dynamics, and other variables related to trophic structure, and linking these patterns to potential changes in oceanographic conditions over time in the Pacific and Atlantic Oceans. Fur seal photo by A. Orr.


