McGinnis Lab Home
Page
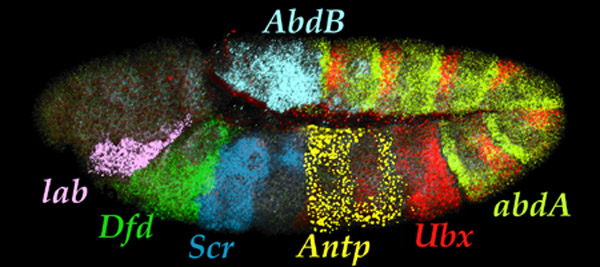
| Seven Hox transcript patterns
in a single embryo. |
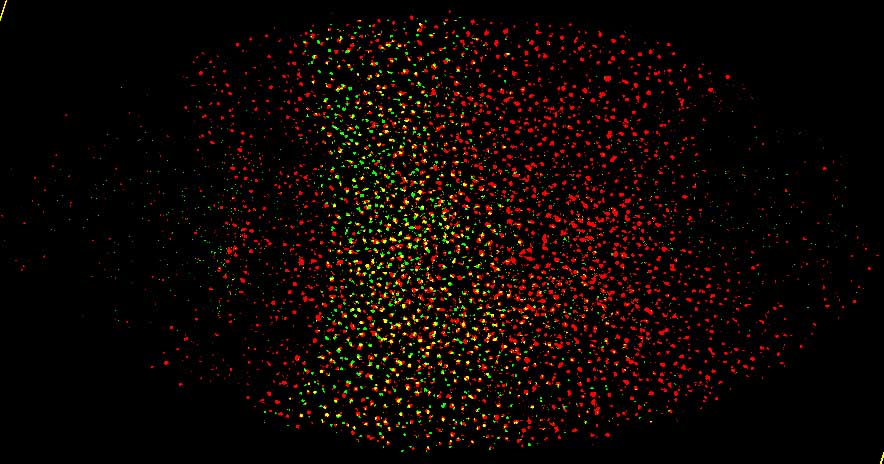
microRNA-10 transcript pattern (red) in embryo |
 |
 |
 |
 |
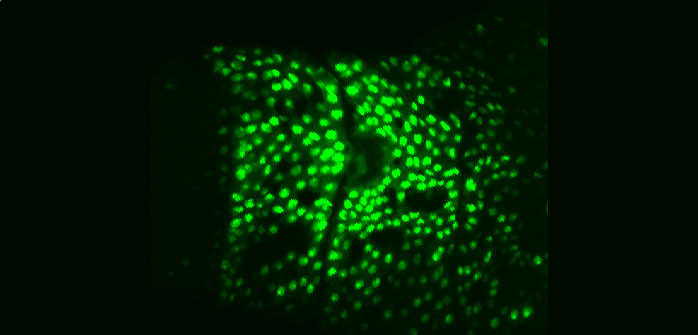
| Aseptic wound response pathway
in embryos |
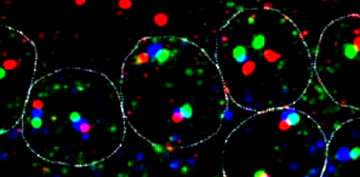
Transcription
units: Dfd, Scr, and ftz in nuclei |
 |
William McGinnis
University of California, San Diego
Division of Biology
Section of Cell & Developmental Biology
9500 Gilman Drive
La Jolla, CA 92093-0349
(858) 822-0458
|
|
|
Bill McGinnis
received his Ph.D. from UC Berkeley in 1982 and was a Jane Coffin
Childs
postdoctoral fellow at the University of Basel. He was a
Professor at Yale University from 1984 until 1995, then
moved his lab to the University of California, San Diego.
He served as the
Chairman of the Department of Biology from July 1998 - June
1999, as Associate Dean of the Division of Natural Sciences
from July 1, 1999 - June 2000, and has been Dean of the Division of Biological Sciences since 2013.
The
McGinnis lab is located at the University
of California, San Diego,
in La
Jolla, CA. The lab is on the fourth floor of Bonner Hall, on the same floor
as the laboratories of James Posakony, Ethan Bier, Steve Wasserman, Chih-ying Su,
Amy Pasquinelli, and Emily Troeml. Our labs have overlapping
interests,
shared resources, and
fruitful
interactions.
The University and the greater La Jolla area form
a community rich in world class research in life sciences and biotechnology.
We are within walking distance of the Salk
Institute, the Scripps
Research Institute, the Burnham Research
Institute and the Scripps
Institution of Oceanography.
Graduate students in the Division of Biological
Sciences at UCSD enjoy the opportunity to rotate through and to choose labs
at
both the UCSD campus and the Salk Institute. Return
to top of
page / People / Publications |
Research
Interests:
1. We are exploring a novel epidermal wound response pathway
in Drosophila. This pathway
can be activated by loss of all Hox functions in a body segment, or by aseptic
puncture
wounds. We have aseptic wound response genes, and the cis-regulatory regions
that activate reporter gene expression in cells adjacent to wounds (see picture
above). The activation of some of these genes in response to breaks in the integument
requires
the Grainy head transcription
factor.
Grainy head protein homologs
are
conspicuously
expressed
in
the
barrier epithelium of both flies and mammals, and required for the formation
and repair of an impermeable "skin" whether that skin is make of keratinocytes
or cuticle. (Mace et al. 2005). We have recently identified many novel genes that regulate the transcription response to epidermal puncture wounds in Drosophila. The functions of these genes indicate that hydrogen peroxide is one signal required for activating regeneration genes around epidermal wounds, and Flotillin-2 is inhibiting the spread of wound signals in Drosophila epidermal cells. (Juarez et al., 2011). Our study of mutants in a fungal (Neurospora) grainy head-like gene suggest that an ancestral grainy head like function was to regulate cell wall composition and remodeling, and in animals, that function was co-opted to control the formation of an apical extracellular matrix and cellular junctions that provide epithelial barrier functions. (Paré et al., 2012). The activation of multiple signaling pathways are required to induce wound dependent transcription, one pathway that is induced is a receptor tyrosine kinase pathway, but it is likely that at least two other pathways are required. It is still unknown what wound-induced ligands (presumably damage associated molecular patterns, or DAMPs) diffuse from the wound to activate wound response transcription. We also seek to understand the kinetics of the spread of wound induced transcription, and are using live transcriptional reporters to study this phenomenon.
2. We have developed methods (multicolor in situ hybridization coupled with high resolution microscopy) to count mRNAs in the cytoplasm
embryonic cells, at the same time as detecting transcription per se of the
same gene in nuclei,
in order to understand gene regulation of animal developmental pathways
at the single cell level.
For
a recent
papers on this, see Pare et al. (2009). and Arvey et al. (2010). We are following up this research with multicolor in situ hybridization experiments to study the function of long-noncoding RNAs that regulated the expression patterns of Hox genes in developing Drosophila.
Return to top of
page / People
/ Publications




